Simple plotting from the command line
December 18, 2013 at 10:53 PM by Dr. Drang
The problem with making graphs in Matplotlib, Gnuplot, and even Numbers, is that it takes too long to get something that looks halfway decent1 when I’m not in “graph-making mode.” I’ll be working on a problem and realize that a quick visualization of a function or some data would help my understanding. I don’t want my train of thought derailed by the context shift that comes with starting up a program and fiddling with it—I just want a reasonable-looking graph now. So I wrote a couple of little scripts to make plots from the command line.
The scripts are in this GitHub repository, and I’ll quote its README to explain how they’re used.
Two simple scripts for creating plots from the command line: one for plotting functions and the other for plotting points. The scripts require the NumPy and matplotlib modules.
The graphs produced aren’t intended to be of publication quality. The goal is to make readable graphs very quickly.
Functions
For these, everything is given on the command line.
plot 'function(s)' minx maxx [miny maxy]
Three arguments are required: the function itself, which would normally be enclosed in quotes to avoid problems with shell interpretation; the minimum x value; and the maximum x value. The last two arguments, the minimum and maximum y values are optional—if they aren’t given, the script will figure them out. A common reason to specify the y limits is if the function “blows up” between the x limits and the extreme y values make the rest of the graph look flat.
The function can include any Python expression or function, including those in the NumPy library. There’s no need to use a numpy prefix.
More than one function can be specified; separate them with a semicolon. Thus, something like
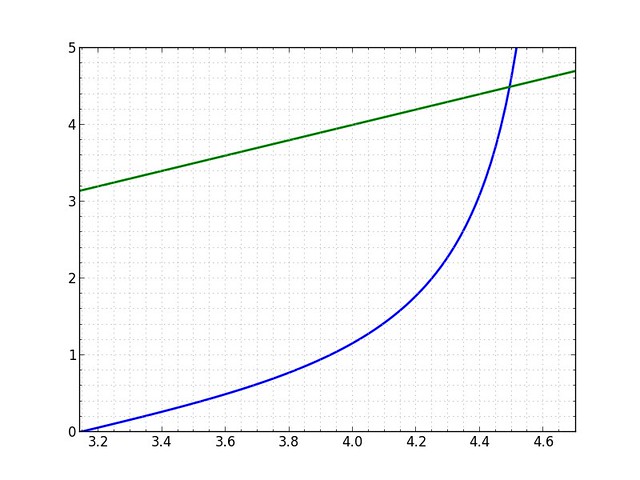
plot 'tan(x); x' pi 3*pi/2-.01 0 5
can be used to find the intersection of the two given functions. This example also shows that you can use expressions in the limits.
The result of plot is an 800×600 PNG image that’s sent to standard output. Normally, this would be redirected to a file,
plot 'tan(x); x' pi 3*pi/2-.01 0 5 > plot.png
and viewed in whatever graphics program the user prefers. No controls for tick marks or grid spacing are provided—this is quick and dirty plotting.
Points
The script that plots lists of points, pplot, gets its data from standard input. On the Mac, it would be common to pass data in from the clipboard this way:
pbpaste | pplot
As with plot, pplot produces an 800×600 PNG image that’s sent to standard output, so one would normally redirect this to a file:
pbpaste | pplot > plot.png
Each line of the input data represents one (x, y) point. The x and y values can be separated by tabs, spaces, or commas.
Normally, pplot plots the points only. If you want to connect the points with a line, use the -l option:
pbpaste | pplot -l > plot.png
Here’s plot:
python:
1: #!/usr/bin/python
2:
3: from __future__ import division
4: from numpy import *
5: import matplotlib.pyplot as plt
6: import sys
7:
8: fcns = sys.argv[1].split(';')
9: minx = eval(sys.argv[2])
10: maxx = eval(sys.argv[3])
11: plt.xlim(minx, maxx)
12: try:
13: miny = float(sys.argv[4])
14: maxy = float(sys.argv[5])
15: plt.ylim(miny, maxy)
16: except IndexError:
17: pass
18:
19: x = linspace(minx, maxx, 100)
20: for i, f in enumerate(fcns):
21: y = eval(f)
22: plt.plot(x, y, "-", linewidth=2)
23: plt.minorticks_on()
24: plt.grid(which='both', color='#aaaaaa')
25: plt.savefig(sys.stdout, format='png')
A few things about plot that I’m not proud of:
- No error handling. The
try/exceptclause isn’t really error handling, it’s just a simple way to handle optional arguments. I’m assuming that if I screw up the input, whatever error message Python spits back will help me fix it. - The
evals. I know it’s considered dangerous, but I’m not going to write my own parser. Plus, I think the danger is overblown. I know perfectly well how the program works; why would I pass in a function that wipes my hard drive? - Bringing in every NumPy command with the
importstatement. I’d rather not have the namespace polluted, but I’ll be damned if I’m going to type things likenp.sin(x)every time I need a trig function.
You may be wondering why I used savefig in Line 25 instead of show. I just don’t like the visualizer that show launches—its window is too small, it blocks further input from the command line, and it requires a mouse click to dismiss it. I’d rather just open a PNG file in Preview. If you prefer show, be my guest; just change Line 25 to plt.show().
Here’s pplot:
python:
1: #!/usr/bin/python
2:
3: import matplotlib.pyplot as plt
4: import sys
5: import re
6: import getopt
7:
8: marker = '.'
9: opts, args = getopt.getopt(sys.argv[1:], 'l')
10: for o, a in opts:
11: if o == '-l':
12: marker = '.-'
13:
14: x = []
15: y = []
16: for line in sys.stdin:
17: line = line.strip()
18: vals = re.split(r'\s+|\s*,\s*', line)
19: x.append(vals[0])
20: y.append(vals[1])
21: plt.plot(x, y, marker)
22: plt.minorticks_on()
23: plt.grid(which='both', color='#aaaaaa')
24: plt.savefig(sys.stdout, format='png')
As you can see in Line 18, the x and y values can be separated by any amount of whitespace or by a comma and whitespace. This makes pplot more flexible in the input it can accept. Like plot, this is a pretty bare-bones script, with no error handling.
An obvious deficiency in pplot is its inability to graph more than one dataset. I thought about extending it, but there are too many ways to make a table with multiple datasets. Once you get to that level of complexity, you’re better off using pandas, which has ways of slicing and handling missing data that a simple program like pplot couldn’t hope to include.
-
Actually, “halfway decent” is beyond Numbers. Its graphs are acceptable, I suppose, for business graphics, where pie charts are considered hot shit, but totally wrong for science and engineering. ↩